Because most NCBI/GenBank sequences are submitted by external researchers using specimens from our collections—and because we are not typically included as co-authors on those submissions and therefore cannot correct or update the sequence data if new information arises—it is important to establish reciprocal links between our Specify records and the corresponding NCBI/GenBank entries. Doing so ensures that researchers can readily see any updates to taxonomic determinations or other relevant information. Along with having clear instructions for lodging sequences as part of any tissue gift/loan agreement this ensures clear attribution and advocacy for collections providing the material as well as making the linkage between specimen voucher, tissue sample and sequence evident to any downstream users of the sequence.
From within Specify, you can create a weblink field that will allow you to use a sequence Genbank number in a field (typically the Genbank number field in the DNA sequence table) to access that sequence on the web.
You can find out how to create a weblink field here: https://discourse.specifysoftware.org/t/editing-web-links-in-specify-7/1559
You can see an example of this functionality from this record in the KU Ichthyology tissue collection by clicking on the DNA button near the bottom of the form - https://ichthyology.specify.ku.edu/specify/bycatalog/KUIT/10/
In order for this to work you will need to have guest access to your collection active as described here: https://discourse.specifysoftware.org/t/enable-anonymous-guest-access-in-specify-7/992
There are two methods of creating reciprocal links in Genbank:
- Structured voucher
- Linkout
Structured voucher
The “/specimen voucher” field in a Genbank nucleotide submission can be used to enter structured voucher information that will link directly to your collection. Information is structured as a Darwin Core Triplet of Institution Code: Collection Code: Catalog number. You then need to provide your information to Genbank to structure the URL That will then link to your collection.
You can see an example of this here: https://www.ncbi.nlm.nih.gov/nuccore/KC831357.
You can register your collection and provide additional information by emailing gb-admin@ncbi.nlm.nih.gov.
More details can be found here: https://www.ncbi.nlm.nih.gov/biocollections/docs/faq/.
Linkout
LinkOut is a service that allows organizations external to NCBI to add and update links to their own resources from individual records in NCBI databases. This service provides visitors with convenient access to outside resources that are intended to extend, clarify, and supplement information found in NCBI databases. This information is displayed in a left-hand menu of the sequence record in Genbank. See this example: https://www.ncbi.nlm.nih.gov/nuccore/KC831357.
Complete documentation on how to utilize this service can be found here: https://www.ncbi.nlm.nih.gov/projects/linkout/ and https://www.ncbi.nlm.nih.gov/books/NBK3802/
Essentially the process consists of multiple steps:
-
Register your collection with the Linkout service by emailing linkout@ncbi.nlm.nih.gov and providing the following information:
-
Name, email address, and phone number of a contact person in your organization.
-
The scope of your resource, including the URL of the resource. If a username and password are required to access the resource, please include a temporary username and password that the LinkOut team can use to evaluate the resource. Also, please describe the type of NCBI database records to which you would like to apply links and provide a couple URL examples of your database records and their corresponding NCBI database records.
-
Describe any restrictions on access to the resource.
-
-
Set up an identity file called providerinfo.xml which identifies your provider information:
- Provider ID – provided by NCBI Linkout
- Name of resource
- Name abbreviation of resource
- Subject type – should be “herbarium/museum collections”
- URL – your collection webpage URL (NOT Specify link)
- Brief – description of resource
Here is an example for my tissue collection:
<?xml version="1.0"?> <!DOCTYPE Provider PUBLIC "-//NLM//DTD LinkOut 1.0//EN" "https://www.ncbi.nlm.nih.gov/projects/linkout/doc/LinkOut.dtd"> <Provider> <ProviderId>9646</ProviderId> <Name>KU Biodiversity Institute</Name> <NameAbbr>kubioinst</NameAbbr> <SubjectType>herbarium/museum collections</SubjectType> <Url>https://biodiversity.ku.edu/Ichthyology/collections</Url> <Brief>University of Kansas Biodiversity Institute Ichthyology Tissue Collection</Brief> </Provider> -
Resource CSV File. This file describes each link between a Specify record and its corresponding sequence(s) and needs to include the following information with column headers:
- PrID – provider ID
- DB – Genbank database
- Genbank – Genbank number
- URL – Specify URL to link to the Specify record
- IconURL – this can be left blank
- Cat # - Catalog number related to the sequence (this is what will show up in the left-hand menu of the sequence record)
- URL Name – this can be left blank
- SubjectType – this can be left blank
- Attribute – this can be left blank
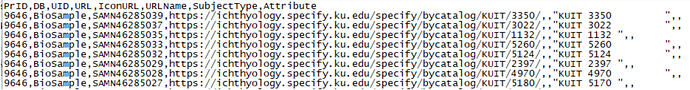
Each sequence should be on a new line and each field should be separated with a comma (including blank fields) as below:
This can also be done for SRA sequences but needs to link to the BioSample record:
-
You then need to set up an FTP file transfer for FTP-private.ncbi.nlm.nih.gov. I use WinSCP (https://winscp.net/eng/index.php). You will be provided with a username and password by NCBI to FTP your files. This needs to include the above providerinfo.xml and your resource CSV file(s). I typically create a new file each year and just add any new sequences to that file and reupload it.